Research Overview
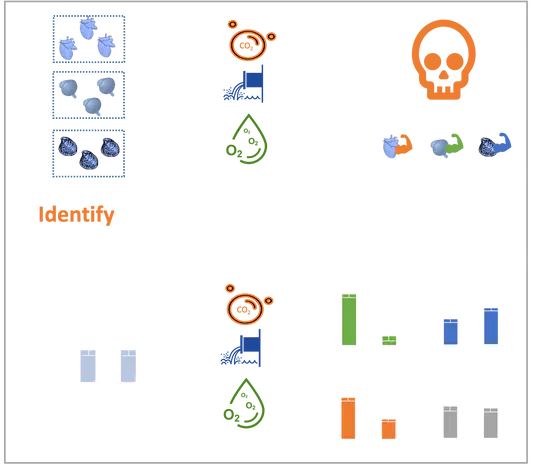
Coastal ecosystems face a complex of stressors that span multiple temporal and spatial scales, from long-term global ocean change to localized episodes of coastal acidification, and marine species experience these multiple stressors simultaneously. Understanding how marine populations will evolve in response to environmental change requires investigating the synergistic impacts of multiple stressors across all life stages.
Our research is divided into two foundational pillars:
- Understanding how natural and anthropogenic processes affect the evolution of marine populations through the lens of larval dispersal.
- We combine laboratory multi-stressor larval exposure experiments with genomic surveys of natural populations, analyzing patterns of selection and migration in a geographic context using landscape (or seascape) genomic models.
- Developing genomic tools to ask evolutionary questions
- We develop both laboratory and bioinformatic methods to facilitate the use of next-generation sequencing in non-model species.
Next-generation sequencing

The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
.js-id-EecSeq
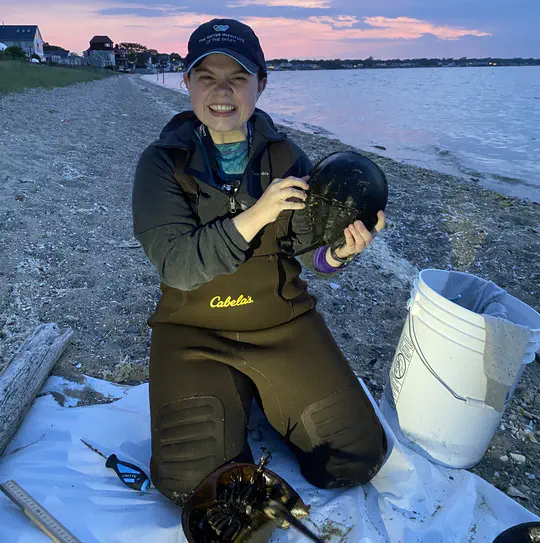
The Atlantic horseshoe crab, Limulus polyphemus, is a commercially, ecologically, and economically important species. Current management practices of this species may not be well informed enough to avoid jeopardizing the future health of these animals.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
dDocent is simple bash wrapper to QC, assemble, map, and call SNPs from almost any kind of RAD sequencing. If you have a reference already, dDocent can be used to call SNPs from almost any type of NGS data set.
The eastern oyster, Crassostrea virginica, is a valuable fishery and aquaculture species that provides critical services as an ecosystem engineer. Oysters have a life-history that promotes high genetic diversity and gene flow while also occupying a wide range of habitats in variable coastal environments from the southern Gulf of Mexico to the southern waters of Atlantic Canada.
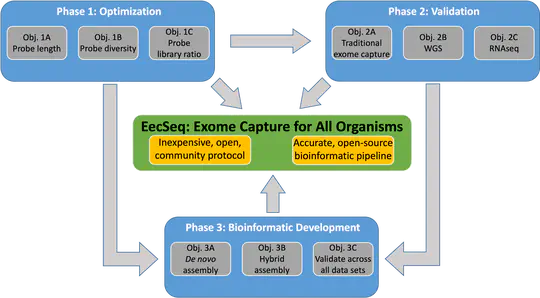
To develop EecSeq into an exome capture method for any organism, the accompanying bioinformatics pipeline needs to be capable of de novo assembly of exon loci directly from captured genomic reads.
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change.
An example of linking directly to an external project website using
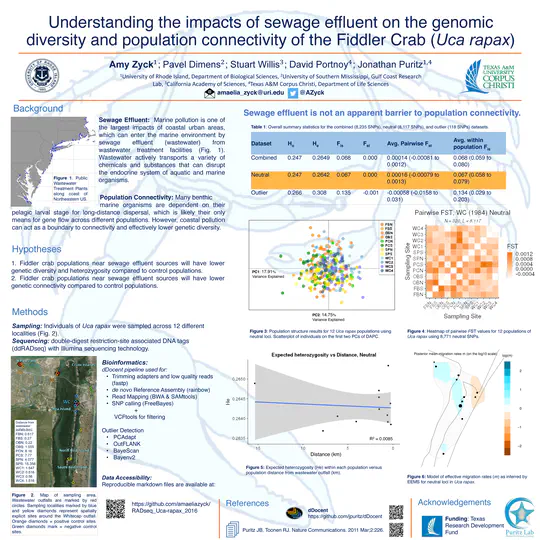
external_link.Coastal urban areas, home to over 50% of the population in the USA and over 60% of the population of the world, are a major source of marine pollution in coastal environments.
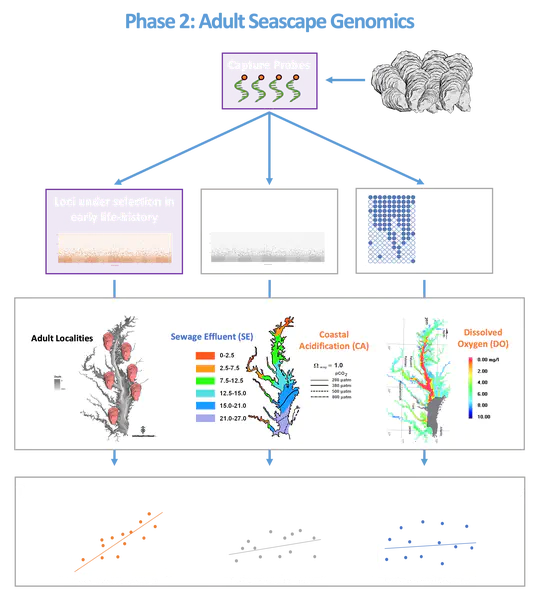
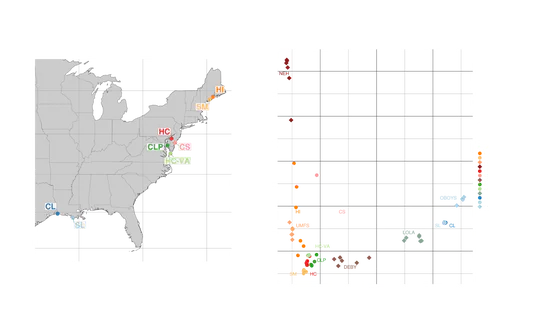
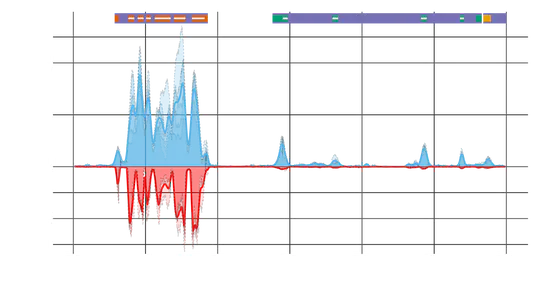
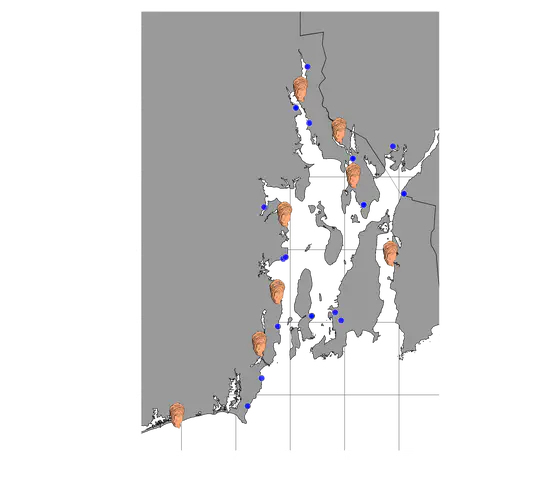
Adult oysters were also collected from various sites in Narragansett Bay in 2017 and 2020. Tissue samples were taken, and DNA was extracted for sequencing. Environmental data like temperature, salinity, pH, dissolved oxygen, and chlorophyll-a concentrations were recorded at each site, and the influence of nearby sewage effluent was assessed.Probes created from the larval oyster mRNA were used to capture specific gene regions in the adult oysters’ DNA. This allowed researchers to study the genetic response to the environmental stressors: coastal acidification and sewage effluent. The objectives of the study were to identify genetic loci under selection, analyze how environmental factors influence these loci, and understand the genetic adaptation from larval to adult stages in response to coastal stressors.
The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
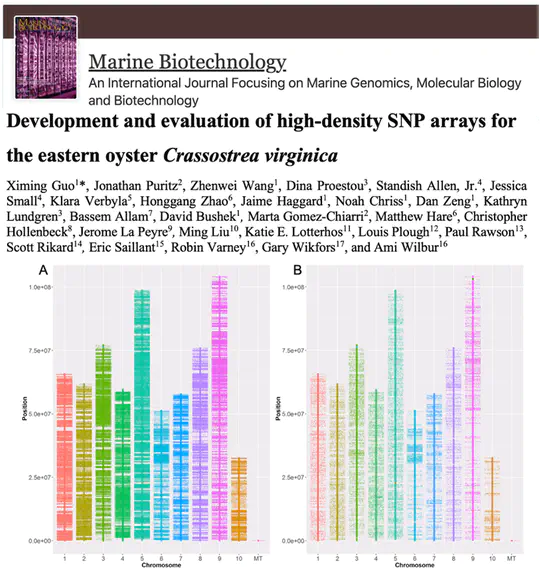
The eastern oyster, a crucial species for US aquaculture, benefits from improved breeding techniques. The Eastern Oyster Breeding Consortium created two SNP arrays to aid in genetic studies. One array, with 566K SNPs, is for screening, while the other, with 66K SNPs, is for breeders’ use.
During the summer of 2017, preliminary exposure trials were conducted using larvae from the eastern oyster (Crassostrea virginica). Wild adult broodstock were collected from Ipswich, MA and Barnstable, MA, and brought into the lab and conditioned for several weeks.
The eastern oyster, Crassostrea virginica, now has its first complete chromosome-level genome assembly. Released and annotated in 2017, this genome is over 97% complete and serves as a vital resource for studying how mollusks adapt to changing environments and for improving selective breeding in aquaculture.
Bioinformatics

As genomic scale data sets become standard in molecular ecology, understanding how to efficiently and accurately process raw data is critical for accuracy in downstream population-level analysis.
.js-id-EecSeqBio
The Atlantic horseshoe crab, Limulus polyphemus, is a commercially, ecologically, and economically important species. Current management practices of this species may not be well informed enough to avoid jeopardizing the future health of these animals.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
dDocent is simple bash wrapper to QC, assemble, map, and call SNPs from almost any kind of RAD sequencing. If you have a reference already, dDocent can be used to call SNPs from almost any type of NGS data set.
The eastern oyster, Crassostrea virginica, is a valuable fishery and aquaculture species that provides critical services as an ecosystem engineer. Oysters have a life-history that promotes high genetic diversity and gene flow while also occupying a wide range of habitats in variable coastal environments from the southern Gulf of Mexico to the southern waters of Atlantic Canada.
To develop EecSeq into an exome capture method for any organism, the accompanying bioinformatics pipeline needs to be capable of de novo assembly of exon loci directly from captured genomic reads.
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change.
An example of linking directly to an external project website using
external_link.Coastal urban areas, home to over 50% of the population in the USA and over 60% of the population of the world, are a major source of marine pollution in coastal environments.
Adult oysters were also collected from various sites in Narragansett Bay in 2017 and 2020. Tissue samples were taken, and DNA was extracted for sequencing. Environmental data like temperature, salinity, pH, dissolved oxygen, and chlorophyll-a concentrations were recorded at each site, and the influence of nearby sewage effluent was assessed.Probes created from the larval oyster mRNA were used to capture specific gene regions in the adult oysters’ DNA. This allowed researchers to study the genetic response to the environmental stressors: coastal acidification and sewage effluent. The objectives of the study were to identify genetic loci under selection, analyze how environmental factors influence these loci, and understand the genetic adaptation from larval to adult stages in response to coastal stressors.
The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
The eastern oyster, a crucial species for US aquaculture, benefits from improved breeding techniques. The Eastern Oyster Breeding Consortium created two SNP arrays to aid in genetic studies. One array, with 566K SNPs, is for screening, while the other, with 66K SNPs, is for breeders’ use.
During the summer of 2017, preliminary exposure trials were conducted using larvae from the eastern oyster (Crassostrea virginica). Wild adult broodstock were collected from Ipswich, MA and Barnstable, MA, and brought into the lab and conditioned for several weeks.
The eastern oyster, Crassostrea virginica, now has its first complete chromosome-level genome assembly. Released and annotated in 2017, this genome is over 97% complete and serves as a vital resource for studying how mollusks adapt to changing environments and for improving selective breeding in aquaculture.
Population Connectivity

Life history traits, such as location of fertilization and mode of larval development, have long been thought to influence the evolutionary process of marine organisms, from fine-scale genetic structure to the tempo and mode of speciation.
.js-id-FiddlerCrab
The Atlantic horseshoe crab, Limulus polyphemus, is a commercially, ecologically, and economically important species. Current management practices of this species may not be well informed enough to avoid jeopardizing the future health of these animals.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
dDocent is simple bash wrapper to QC, assemble, map, and call SNPs from almost any kind of RAD sequencing. If you have a reference already, dDocent can be used to call SNPs from almost any type of NGS data set.
The eastern oyster, Crassostrea virginica, is a valuable fishery and aquaculture species that provides critical services as an ecosystem engineer. Oysters have a life-history that promotes high genetic diversity and gene flow while also occupying a wide range of habitats in variable coastal environments from the southern Gulf of Mexico to the southern waters of Atlantic Canada.
To develop EecSeq into an exome capture method for any organism, the accompanying bioinformatics pipeline needs to be capable of de novo assembly of exon loci directly from captured genomic reads.
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change.
An example of linking directly to an external project website using
external_link.Coastal urban areas, home to over 50% of the population in the USA and over 60% of the population of the world, are a major source of marine pollution in coastal environments.
Adult oysters were also collected from various sites in Narragansett Bay in 2017 and 2020. Tissue samples were taken, and DNA was extracted for sequencing. Environmental data like temperature, salinity, pH, dissolved oxygen, and chlorophyll-a concentrations were recorded at each site, and the influence of nearby sewage effluent was assessed.Probes created from the larval oyster mRNA were used to capture specific gene regions in the adult oysters’ DNA. This allowed researchers to study the genetic response to the environmental stressors: coastal acidification and sewage effluent. The objectives of the study were to identify genetic loci under selection, analyze how environmental factors influence these loci, and understand the genetic adaptation from larval to adult stages in response to coastal stressors.
The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
The eastern oyster, a crucial species for US aquaculture, benefits from improved breeding techniques. The Eastern Oyster Breeding Consortium created two SNP arrays to aid in genetic studies. One array, with 566K SNPs, is for screening, while the other, with 66K SNPs, is for breeders’ use.
During the summer of 2017, preliminary exposure trials were conducted using larvae from the eastern oyster (Crassostrea virginica). Wild adult broodstock were collected from Ipswich, MA and Barnstable, MA, and brought into the lab and conditioned for several weeks.
The eastern oyster, Crassostrea virginica, now has its first complete chromosome-level genome assembly. Released and annotated in 2017, this genome is over 97% complete and serves as a vital resource for studying how mollusks adapt to changing environments and for improving selective breeding in aquaculture.
Selection

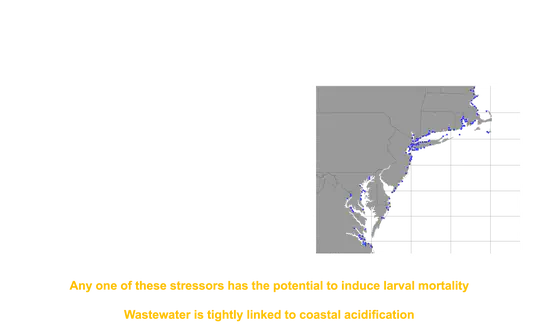
The environmental impacts of an ever-growing human population cross multiple spatial and temporal boundaries, and marine organisms experience all of these stressors simultaneously, increasing the need for research on synergistic effects.
.js-id-Coastal_Stressors
The Atlantic horseshoe crab, Limulus polyphemus, is a commercially, ecologically, and economically important species. Current management practices of this species may not be well informed enough to avoid jeopardizing the future health of these animals.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
dDocent is simple bash wrapper to QC, assemble, map, and call SNPs from almost any kind of RAD sequencing. If you have a reference already, dDocent can be used to call SNPs from almost any type of NGS data set.
The eastern oyster, Crassostrea virginica, is a valuable fishery and aquaculture species that provides critical services as an ecosystem engineer. Oysters have a life-history that promotes high genetic diversity and gene flow while also occupying a wide range of habitats in variable coastal environments from the southern Gulf of Mexico to the southern waters of Atlantic Canada.
To develop EecSeq into an exome capture method for any organism, the accompanying bioinformatics pipeline needs to be capable of de novo assembly of exon loci directly from captured genomic reads.
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change.
An example of linking directly to an external project website using
external_link.Coastal urban areas, home to over 50% of the population in the USA and over 60% of the population of the world, are a major source of marine pollution in coastal environments.
Adult oysters were also collected from various sites in Narragansett Bay in 2017 and 2020. Tissue samples were taken, and DNA was extracted for sequencing. Environmental data like temperature, salinity, pH, dissolved oxygen, and chlorophyll-a concentrations were recorded at each site, and the influence of nearby sewage effluent was assessed.Probes created from the larval oyster mRNA were used to capture specific gene regions in the adult oysters’ DNA. This allowed researchers to study the genetic response to the environmental stressors: coastal acidification and sewage effluent. The objectives of the study were to identify genetic loci under selection, analyze how environmental factors influence these loci, and understand the genetic adaptation from larval to adult stages in response to coastal stressors.
The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
The eastern oyster, a crucial species for US aquaculture, benefits from improved breeding techniques. The Eastern Oyster Breeding Consortium created two SNP arrays to aid in genetic studies. One array, with 566K SNPs, is for screening, while the other, with 66K SNPs, is for breeders’ use.
During the summer of 2017, preliminary exposure trials were conducted using larvae from the eastern oyster (Crassostrea virginica). Wild adult broodstock were collected from Ipswich, MA and Barnstable, MA, and brought into the lab and conditioned for several weeks.
The eastern oyster, Crassostrea virginica, now has its first complete chromosome-level genome assembly. Released and annotated in 2017, this genome is over 97% complete and serves as a vital resource for studying how mollusks adapt to changing environments and for improving selective breeding in aquaculture.
Seascape Genomics

We determine the role of natural and anthropogenic forces shaping the evolution of marine populations by testing (i) if selective regimes differ and interact across life-history stages and (ii) if the frequencies of both neutral and resistant genotypes correlate to environmental conditions and if adaptive loci partitioned across life stage.
.js-id-Coastal_Stressors_SG
The Atlantic horseshoe crab, Limulus polyphemus, is a commercially, ecologically, and economically important species. Current management practices of this species may not be well informed enough to avoid jeopardizing the future health of these animals.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
Marine species face a complex suite of stressors that span multiple temporal and spatial scales from long-term global ocean change to localized episodes of coastal acidification. The cumulative and concurrent impacts of multiple stressors remain relatively unknown and requires investigating their synergistic impacts across all life stages.
dDocent is simple bash wrapper to QC, assemble, map, and call SNPs from almost any kind of RAD sequencing. If you have a reference already, dDocent can be used to call SNPs from almost any type of NGS data set.
The eastern oyster, Crassostrea virginica, is a valuable fishery and aquaculture species that provides critical services as an ecosystem engineer. Oysters have a life-history that promotes high genetic diversity and gene flow while also occupying a wide range of habitats in variable coastal environments from the southern Gulf of Mexico to the southern waters of Atlantic Canada.
To develop EecSeq into an exome capture method for any organism, the accompanying bioinformatics pipeline needs to be capable of de novo assembly of exon loci directly from captured genomic reads.
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change.
An example of linking directly to an external project website using
external_link.Coastal urban areas, home to over 50% of the population in the USA and over 60% of the population of the world, are a major source of marine pollution in coastal environments.
Adult oysters were also collected from various sites in Narragansett Bay in 2017 and 2020. Tissue samples were taken, and DNA was extracted for sequencing. Environmental data like temperature, salinity, pH, dissolved oxygen, and chlorophyll-a concentrations were recorded at each site, and the influence of nearby sewage effluent was assessed.Probes created from the larval oyster mRNA were used to capture specific gene regions in the adult oysters’ DNA. This allowed researchers to study the genetic response to the environmental stressors: coastal acidification and sewage effluent. The objectives of the study were to identify genetic loci under selection, analyze how environmental factors influence these loci, and understand the genetic adaptation from larval to adult stages in response to coastal stressors.
The advent of next-generation sequencing (NGS) has rapidly transcended population genetics to population genomics. Current research focuses on adopting next-generation sequencing technology and embracing an ever-adapting genomic toolkit to take advantage of this unprecedented amount of genetic data.
The eastern oyster, a crucial species for US aquaculture, benefits from improved breeding techniques. The Eastern Oyster Breeding Consortium created two SNP arrays to aid in genetic studies. One array, with 566K SNPs, is for screening, while the other, with 66K SNPs, is for breeders’ use.
During the summer of 2017, preliminary exposure trials were conducted using larvae from the eastern oyster (Crassostrea virginica). Wild adult broodstock were collected from Ipswich, MA and Barnstable, MA, and brought into the lab and conditioned for several weeks.
The eastern oyster, Crassostrea virginica, now has its first complete chromosome-level genome assembly. Released and annotated in 2017, this genome is over 97% complete and serves as a vital resource for studying how mollusks adapt to changing environments and for improving selective breeding in aquaculture.