Expressed Exome Capture Sequencing
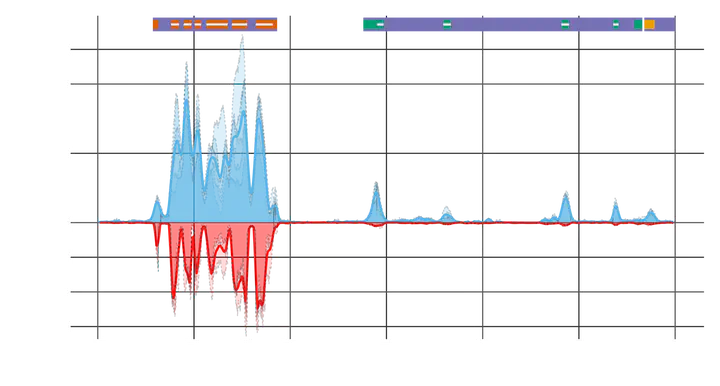
Understanding the interaction between genotype, phenotype, and the environment is one of the greatest challenges in biology. Researchers face a two-fold challenge in experimental design: 1) sampling enough individuals to accurately characterize populations and 2) sequencing the most informative part of the genome, the base pairs that cause phenotypic change. Even with major technological advances in DNA sequencing, it is still too expensive, for most organisms, to sequence the entire genome of multiple individuals. Using probes to target specific locations in the genome has the advantage of focusing sequencing on the functional areas of the genome. However, current methods require preexisting genomic resources for probe design and substantial financial resources for probe synthesis, limiting use to already well-studied organisms and long-term projects. This risky project will develop a method for capturing specific areas of the genome for sequencing with no need for existing genomic resources, removing the time and cost of probe development. This new method will enable the rapid and cost-effective sequencing of the portions of the genome involved in adaptation and will provide unprecedented capacity to detect selection in captive and wild populations. If successful, this new method will enable the assessment of rapid adaptation to short-term ecological disasters and long-term climate change, directly assisting with successful mitigation, conservation, and restoration efforts.
The over-arching goal of the proposed research is to develop Expressed Exome Capture Sequencing (EecSeq) into a cost and time efficient method of exome capture for any organism. To do this, the project takes a three phased approach, (1) focusing on laboratory protocol optimization and improvement, (2) validation with whole genome sequencing (WGS), traditional exome capture, and RNAseq, and (3) the development of an open source, reproducible bioinformatics pipeline, including de novo assembly. Phase one will optimize three key elements in the EecSeq protocol: probe and insert length, probe and capture pool diversity, and the hybridization process. Optimizing all three parameters will maximize the number of sequenced exomic basepairs and on-target reads, increasing the number of individuals that can be sequenced simultaneously while greatly reducing costs. The second phase of the project will use two independent approaches to validate EecSeq genotypes, including a comparison of EecSeq to traditional exome capture, a set of reference individuals with WGS data. The final phase will leverage the chromosome-level assembly of the eastern oyster genome along with results from both previous phases to develop two complementing de novo assembly methods for EecSeq: one utilizing captured genomic reads and a hybrid method that will utilize sequences from the cDNA probes (when sequenced) and captured genomic reads. All experiments, data analysis, and presentation will take place in a completely open and reproducible science pipeline, which should lead to an efficient step-by-step laboratory protocol and a de novo bioinformatic pipeline for EecSeq that incorporates locus assembly and annotation for any organism.
